Quality
Quality traits for salmon
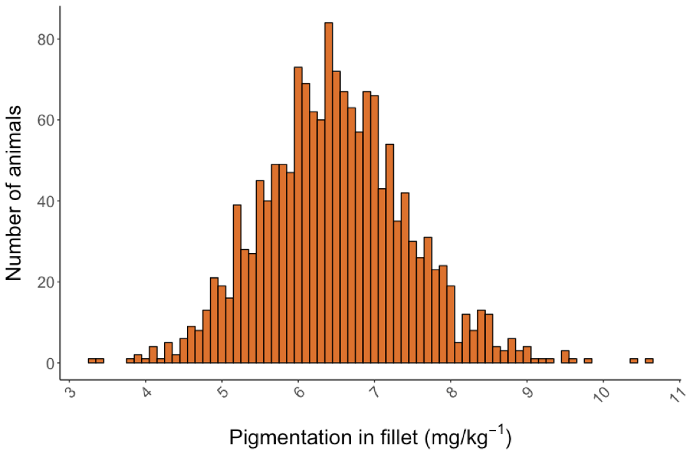
Fillet Pigmentation is one of the most interesting heritable quality traits for Atlantic salmon. The phenotype has been measured both manually as a SalmoFan score and with the help of spectroscopic devices. A Near Infrared (NIR) measurement has also been used, either as a point measurement within the Norwegian Quality Cut (NQC) or as an average measurement of the entire fillet. Within the salmon population, there is a broad variety of the amount of astaxanthin per kilo detected in each fillet. Therefore this trait is reasonable to improve through selective breeding (Figure 1). Fillet pigmentation is recorded annually on the full-sibs of our breeding candidates, during our test slaughter.
SNP Markers
For each genetic trait the SNP markers that are evenly distributed over the entire salmon genome, contributes differently. To map the contribution of each marker towards fillet pigmentation, the adipose fins are sampled for genotyping and connected to the phenotype of each animal. From there it is estimated through a genome-wide association study (GWAS). For SalmoBreed strain of the year-class 2015, 1,698 fish were genotyped with a custom-made Affymetrix SNP array, NOFSAL03, consisting of more than 50K SNP markers.
A GWAS was also performed in order to link their individual measures of mg/kg-1 astaxanthin on the whole fillet to its genome expressions (Figure 2). The Manhattan plot from this GWAS clearly shows that one QTL affects the pigmentation level in a larger degree than the rest of the genome. It also shows that minor components of the rest of the genome contributes to some degree.
Benchmark Genetics offers GS and QTL for pigmentation. Since 2017 these technologies have been implemented to improve the quality of these phenotypes. However it has been part of the SalmoBreed strain’s breeding program since 2005, with the aim to steadily increase the fillet pigmentation of the breeding nucleus. For our GS Quality product we have included GS for production yield and pigmentation, as well as QTL for pigmentation.